Case Study: Replicating Mishra et al. (2020)
Sebastian Funk
2026-04-15
Source:vignettes/mishra-case-study.Rmd
mishra-case-study.RmdIntroduction
This vignette demonstrates how to replicate the analysis from Mishra et al. (2020) of COVID-19 transmission dynamics in South Korea using EpiAwareR. This case study showcases the compositional modelling approach by building a complete epidemiological model from three reusable components.
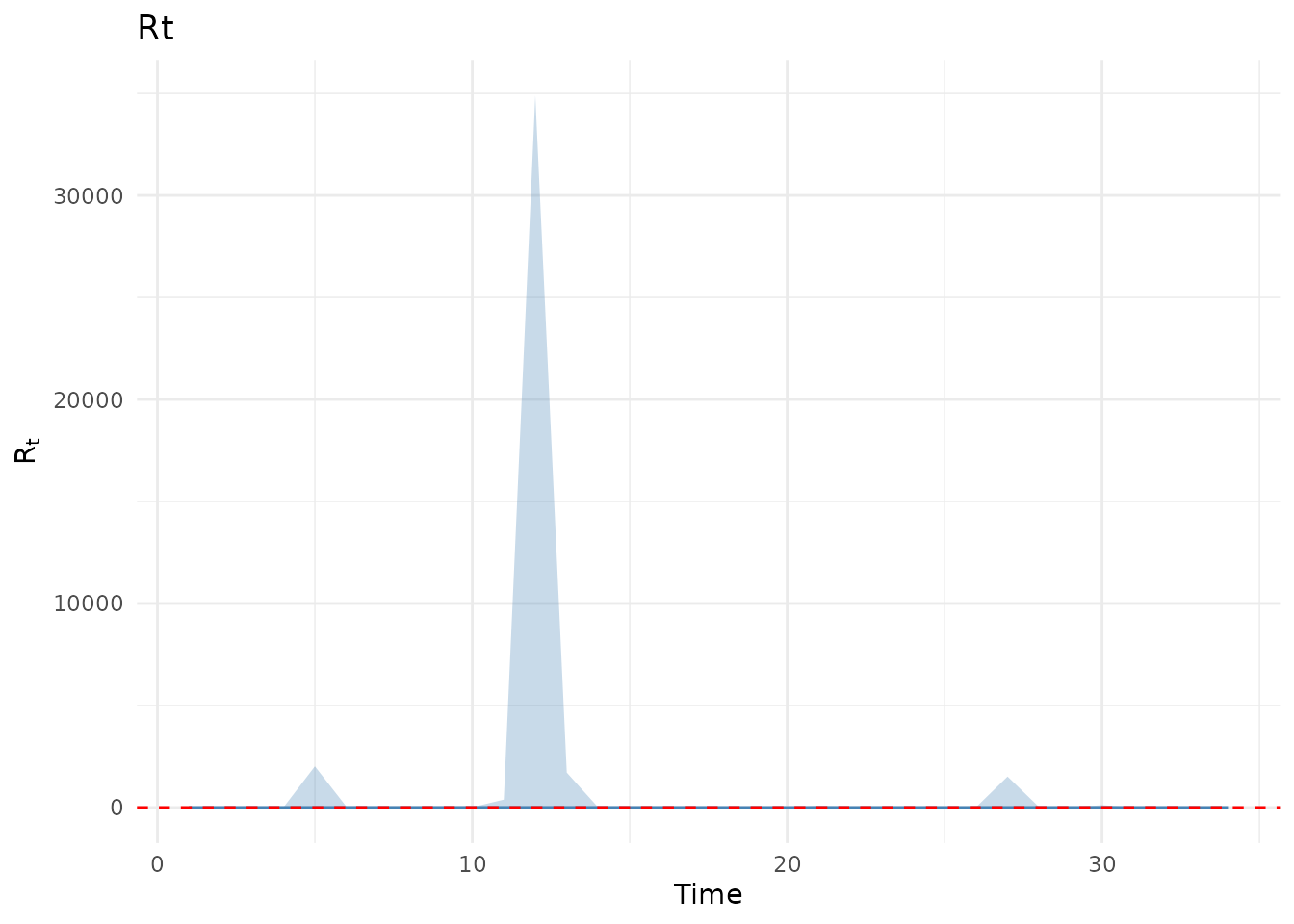
The analysis estimates the time-varying reproduction number () using:
- AR(2) latent process for smooth evolution of log
- Renewal equation for infection dynamics
- Negative binomial observation model for overdispersed case counts
Data Preparation
We use the actual South Korea COVID-19 data from the preprint, which
is included in the package. The data comes from the
covidregionaldata R package and contains daily confirmed
cases from January to July 2020.
# Load South Korea COVID-19 data
data_path <- system.file("extdata", "south_korea_data.csv", package = "EpiAwareR")
south_korea <- read.csv(data_path)
south_korea$date <- as.Date(south_korea$date)
# Prepare full dataset - tspan will select the window
# Column must be named 'y_t' for EpiAware
full_data <- data.frame(
date = south_korea$date,
y_t = south_korea$cases_new
)
# Preview the training window (days 45-80)
# This corresponds to Feb 13 - Mar 19, 2020 (the main epidemic wave)
cat("Training window (tspan 45-80):\n")
#> Training window (tspan 45-80):
print(full_data[45:80, ])
#> date y_t
#> 45 2020-02-13 0
#> 46 2020-02-14 0
#> 47 2020-02-15 0
#> 48 2020-02-16 1
#> 49 2020-02-17 1
#> 50 2020-02-18 1
#> 51 2020-02-19 15
#> 52 2020-02-20 34
#> 53 2020-02-21 75
#> 54 2020-02-22 190
#> 55 2020-02-23 256
#> 56 2020-02-24 161
#> 57 2020-02-25 130
#> 58 2020-02-26 254
#> 59 2020-02-27 449
#> 60 2020-02-28 427
#> 61 2020-02-29 909
#> 62 2020-03-01 595
#> 63 2020-03-02 686
#> 64 2020-03-03 600
#> 65 2020-03-04 516
#> 66 2020-03-05 438
#> 67 2020-03-06 518
#> 68 2020-03-07 483
#> 69 2020-03-08 367
#> 70 2020-03-09 248
#> 71 2020-03-10 131
#> 72 2020-03-11 242
#> 73 2020-03-12 114
#> 74 2020-03-13 110
#> 75 2020-03-14 107
#> 76 2020-03-15 76
#> 77 2020-03-16 74
#> 78 2020-03-17 84
#> 79 2020-03-18 93
#> 80 2020-03-19 152
# Plot the training data
plot(full_data$date[45:80], full_data$y_t[45:80], type = "b",
xlab = "Date", ylab = "Daily Cases",
main = "South Korea COVID-19 Cases (Feb 13 - Mar 19, 2020)")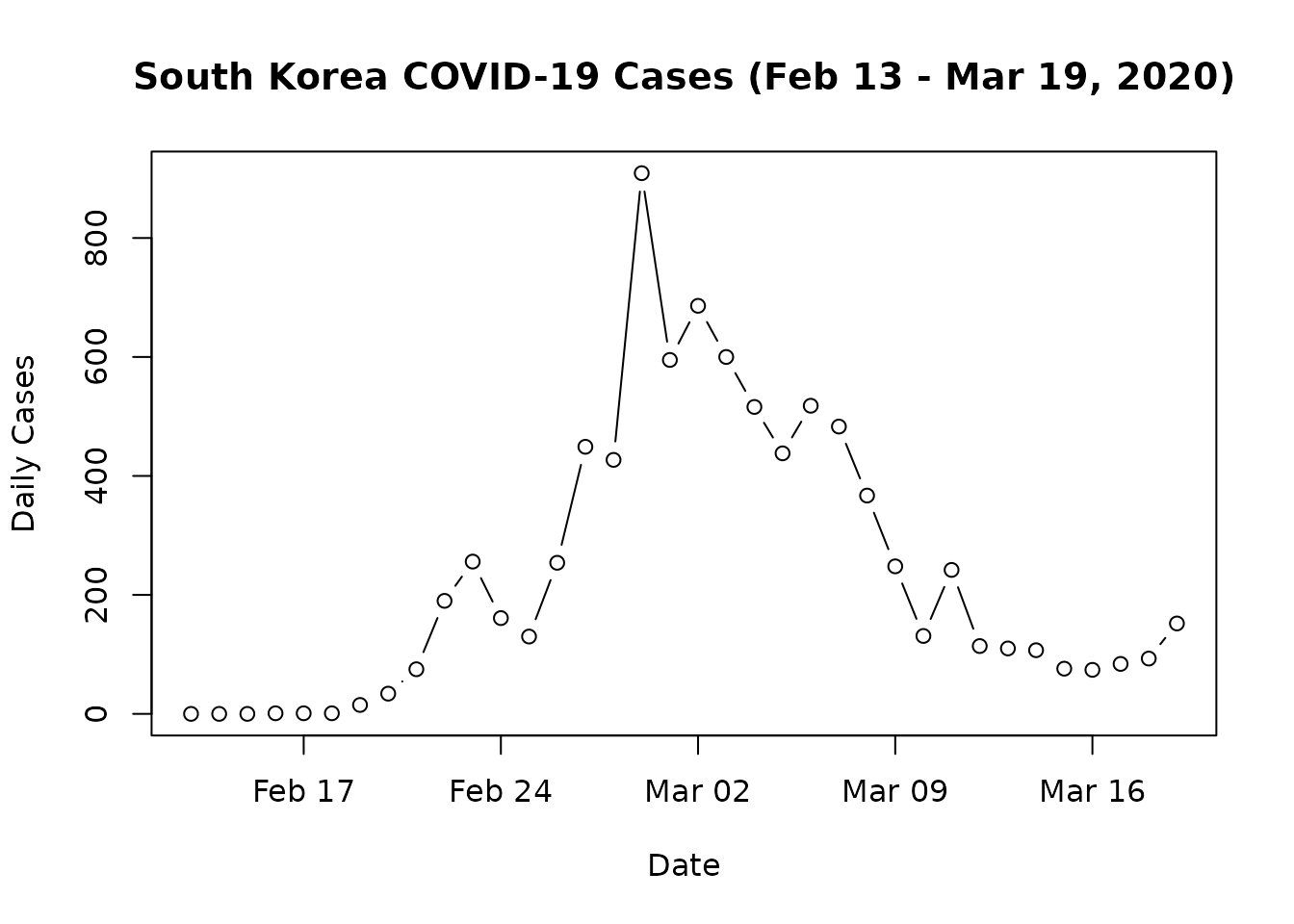
Model Components
1. Latent Process: AR(2) Model
The AR(2) process models the evolution of log over time with temporal autocorrelation:
where .
ar2 <- AR(
order = 2,
damp_priors = list(
truncnorm(0.1, 0.05, 0, 1), # ρ₁: Second-order coefficient
truncnorm(0.8, 0.05, 0, 1) # ρ₂: First-order (strong autocorrelation)
),
init_priors = list(
norm(-1.0, 0.1), # Initial value for lag 1
norm(-1.0, 0.5) # Initial value for lag 2
),
std_prior = halfnorm(0.5) # σ: Innovation standard deviation
)
#> Julia version 1.11.9 at location /opt/hostedtoolcache/julia/1.11.9/x64/bin will be used.
#> Loading setup script for JuliaCall...
#> Finish loading setup script for JuliaCall.
#> EpiAware Julia backend loaded successfully
print(ar2)
#> <EpiAware AR(2) Latent Model>
#> Damping priors: 2
#> Init priors: 2
#> Innovation std prior: specifiedPrior choices (from Mishra et al. 2020):
- First-order damping coefficient ρ₂ centered at 0.8 reflects strong autocorrelation in log Rt
- Second-order damping coefficient ρ₁ centered at 0.1 for smooth dynamics
- Initial values centered at -1 on log scale (Rt ≈ 0.37 initially)
- Innovation std of 0.5 allows flexibility in Rt changes
2. Infection Model: Renewal Equation
The renewal equation models new infections based on past infections and generation time:
where is the discretized generation time distribution.
renewal <- Renewal(
gen_distribution = gamma_dist(6.5, 0.62), # Shape and scale parameters
initialisation_prior = norm(log(1.0), 1.0) # Prior on initial log infections (wide)
)
print(renewal)
#> <EpiAware Renewal Infection Model>
#> Generation distribution: Gamma
#> Initialisation prior: specifiedKey parameters:
- Generation time: Mean ~6.5 days (typical for COVID-19 in early 2020)
- Gamma distribution allows flexible shape
- Initial infections seeded near 1 with wide prior (sd=1.0 on log scale)
3. Observation Model: Negative Binomial
Links latent infections to observed case counts with overdispersion:
where controls overdispersion (clustering).
# The preprint fixes cluster_factor to 0.25
# We approximate this with a tight truncated normal prior
negbin <- NegativeBinomialError(
cluster_factor_prior = truncnorm(0.25, 0.01, 0, 1)
)
print(negbin)
#> <EpiAware Negative Binomial Observation Model>
#> Cluster factor prior: truncated(Normal(mean, sd), lower, upper)Parameterization:
- Cluster factor = (coefficient of variation)
- Fixed to 0.25 (via tight prior) as in the preprint
- More interpretable than directly specifying
Compose and Fit
Create Complete Model
model <- EpiProblem(
epi_model = renewal,
latent_model = ar2,
observation_model = negbin,
tspan = c(45, 80) # Exact tspan from preprint (Feb 13 - Mar 19, 2020)
)
print(model)
#> <EpiAware Epidemiological Model>
#> Time span: 45 to 80
#> Components:
#> - Infection model: epiaware_renewal
#> - Latent model: epiaware_ar
#> - Observation model: epiaware_negbinThe EpiProblem combines components into a joint Bayesian
model with automatic validation. The tspan = c(45, 80)
selects days 45-80 from the full dataset, matching the preprint
exactly.
Bayesian Inference
We use NUTS (No-U-Turn Sampler) with Pathfinder initialization for posterior inference:
results <- fit(
model = model,
data = full_data, # Pass full dataset; tspan selects the window
method = nuts_sampler(
warmup = 1000, # Adaptation iterations
draws = 2000, # Posterior samples per chain (matches preprint)
chains = 4 # Independent chains for convergence assessment
)
)
#> Generating Turing.jl model...
#> Running NUTS sampling...
#> Chains: 4
#> Warmup: 1000
#> Draws: 2000
#> Running Pathfinder initialization...
#> Pathfinder init failed, using default initialization...
#> Processing results...Note: Fitting typically takes 2-5 minutes on modern hardware.
Results
Convergence Diagnostics
print(results)
#> <EpiAware Model Fit>
#>
#> Model:
#> Time span: 45 to 80
#> Infection model: epiaware_renewal
#> Latent model: epiaware_ar
#> Observation model: epiaware_negbin
#>
#> Sampling:
#> Method: NUTS
#> Chains: 4
#> Draws: 2000 (per chain)
#>
#> Convergence:
#> Max Rhat: 1.117
#> Min ESS (bulk): 40
#> Warning: Some parameters have Rhat > 1.1
#> Warning: Some parameters have ESS < 100
#>
#> Use summary() for parameter estimates
#> Use plot() to visualize resultsCheck for:
- Rhat < 1.1: Chains have converged to the same distribution
- ESS > 100: Sufficient effective sample size for inference
- No divergent transitions: NUTS is exploring the posterior efficiently
Parameter Summaries
# Detailed posterior summaries
summary(results)
#> # A tibble: 53 × 10
#> variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 latent.… -0.994 -0.994 0.0995 0.0989 -1.16 -0.829 1.000 10918. 5980.
#> 2 latent.… -0.653 -0.648 0.469 0.460 -1.43 0.107 1.00 6959. 5778.
#> 3 latent.… 0.0841 0.0827 0.0396 0.0420 0.0214 0.152 1.00 5093. 3566.
#> 4 latent.… 0.810 0.810 0.0436 0.0443 0.739 0.880 1.00 4926. 4387.
#> 5 latent.… 0.555 0.547 0.0901 0.0878 0.422 0.717 1.00 3394. 4577.
#> 6 latent.… 0.988 0.982 0.906 0.916 -0.482 2.50 1.00 8434. 5802.
#> 7 latent.… 1.38 1.38 0.898 0.906 -0.0941 2.86 1.00 9138. 5895.
#> 8 latent.… 1.24 1.25 0.920 0.932 -0.269 2.73 1.00 9259. 5526.
#> 9 latent.… 1.18 1.18 0.891 0.879 -0.289 2.63 1.00 7977. 5893.
#> 10 latent.… 2.15 2.14 0.814 0.804 0.813 3.51 1.00 8713. 6120.
#> # ℹ 43 more rowsKey parameters to examine:
- AR damping coefficients (, ): Autocorrelation strength
- Innovation std (): Variability in changes
- Cluster factor: Degree of case count overdispersion
- Initial infections: Epidemic seeding
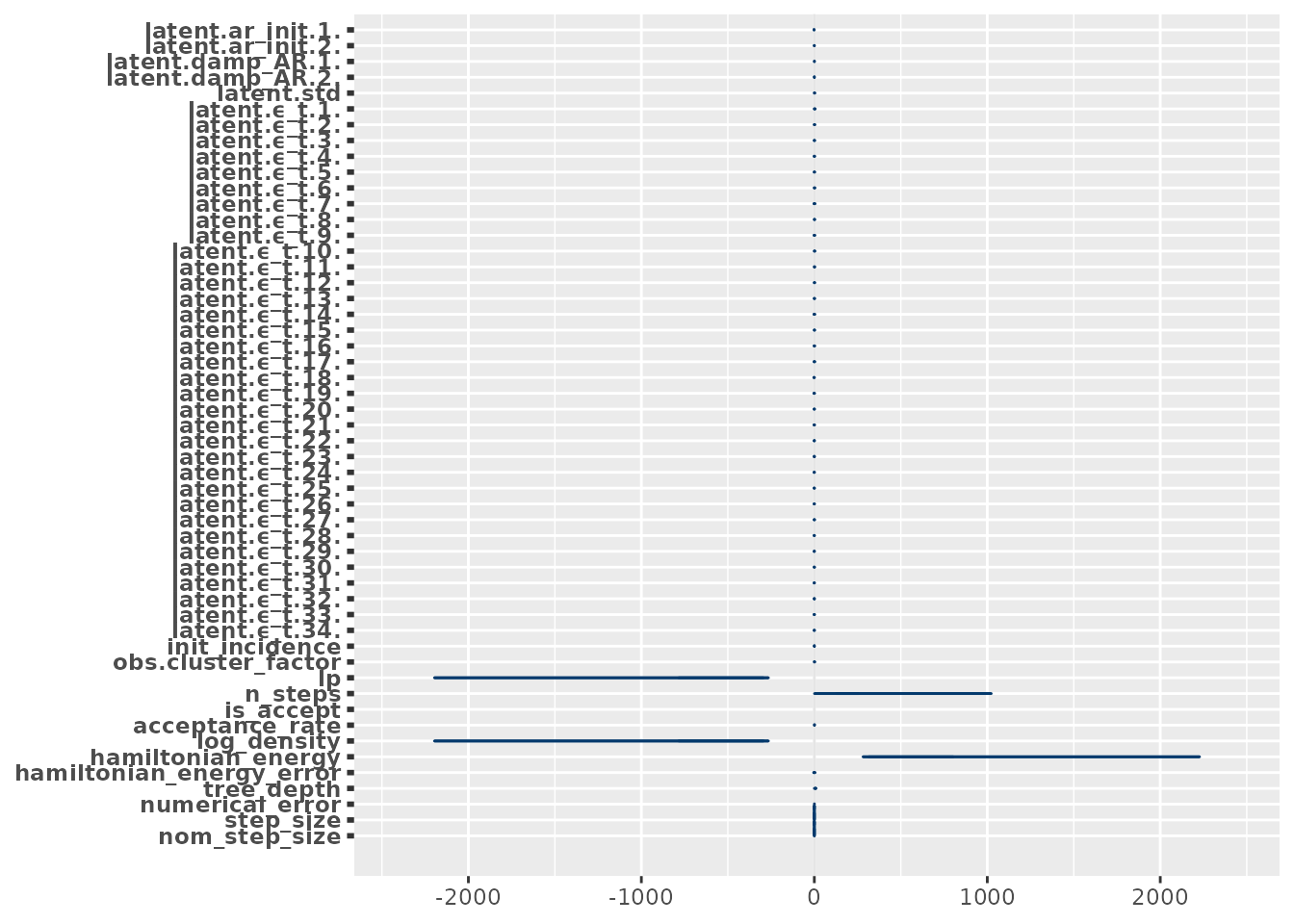
Working with posterior draws
epiaware_fit objects integrate directly with the posterior package. You can
convert the samples to any draws format for use with bayesplot, loo, or
other packages:
library(posterior)
#> This is posterior version 1.7.0
#>
#> Attaching package: 'posterior'
#> The following objects are masked from 'package:stats':
#>
#> mad, sd, var
#> The following objects are masked from 'package:base':
#>
#> %in%, match
# Convert to draws_df for tidy workflows
draws <- as_draws_df(results)
head(draws)
#> # A draws_df: 6 iterations, 1 chains, and 53 variables
#> latent.ar_init.1. latent.ar_init.2. latent.damp_AR.1. latent.damp_AR.2.
#> 1 -1.15 -1.175 0.119 0.81
#> 2 -0.86 0.083 0.077 0.79
#> 3 -1.04 -0.505 0.070 0.81
#> 4 -1.00 -0.187 0.085 0.83
#> 5 -0.94 -0.120 0.129 0.81
#> 6 -0.87 0.316 0.100 0.80
#> latent.std latent.ϵ_t.1. latent.ϵ_t.2. latent.ϵ_t.3.
#> 1 0.44 0.50 2.85 1.67
#> 2 0.49 1.64 0.22 0.52
#> 3 0.60 0.71 1.43 1.45
#> 4 0.57 0.24 0.56 1.46
#> 5 0.60 0.22 0.25 1.76
#> 6 0.53 0.62 0.80 1.21
#> # ... with 45 more variables
#> # ... hidden reserved variables {'.chain', '.iteration', '.draw'}
# Find AR damping and std parameters by pattern
all_vars <- variables(draws)
ar_vars <- grep("damp_AR|latent\\.std", all_vars, value = TRUE)
ar_vars
#> [1] "latent.damp_AR.1." "latent.damp_AR.2." "latent.std"
# Summarise the AR-related parameters
subset_draws(draws, variable = ar_vars) |>
summarise_draws()
#> # A tibble: 3 × 10
#> variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 latent.damp_… 0.0841 0.0827 0.0396 0.0420 0.0214 0.152 1.00 5093. 3566.
#> 2 latent.damp_… 0.810 0.810 0.0436 0.0443 0.739 0.880 1.00 4926. 4387.
#> 3 latent.std 0.555 0.547 0.0901 0.0878 0.422 0.717 1.00 3394. 4577.This also enables bayesplot diagnostics:
library(bayesplot)
# Trace plots for AR damping coefficients
damp_vars <- grep("damp_AR", variables(as_draws_array(results)), value = TRUE)
mcmc_trace(as_draws_array(results), pars = damp_vars)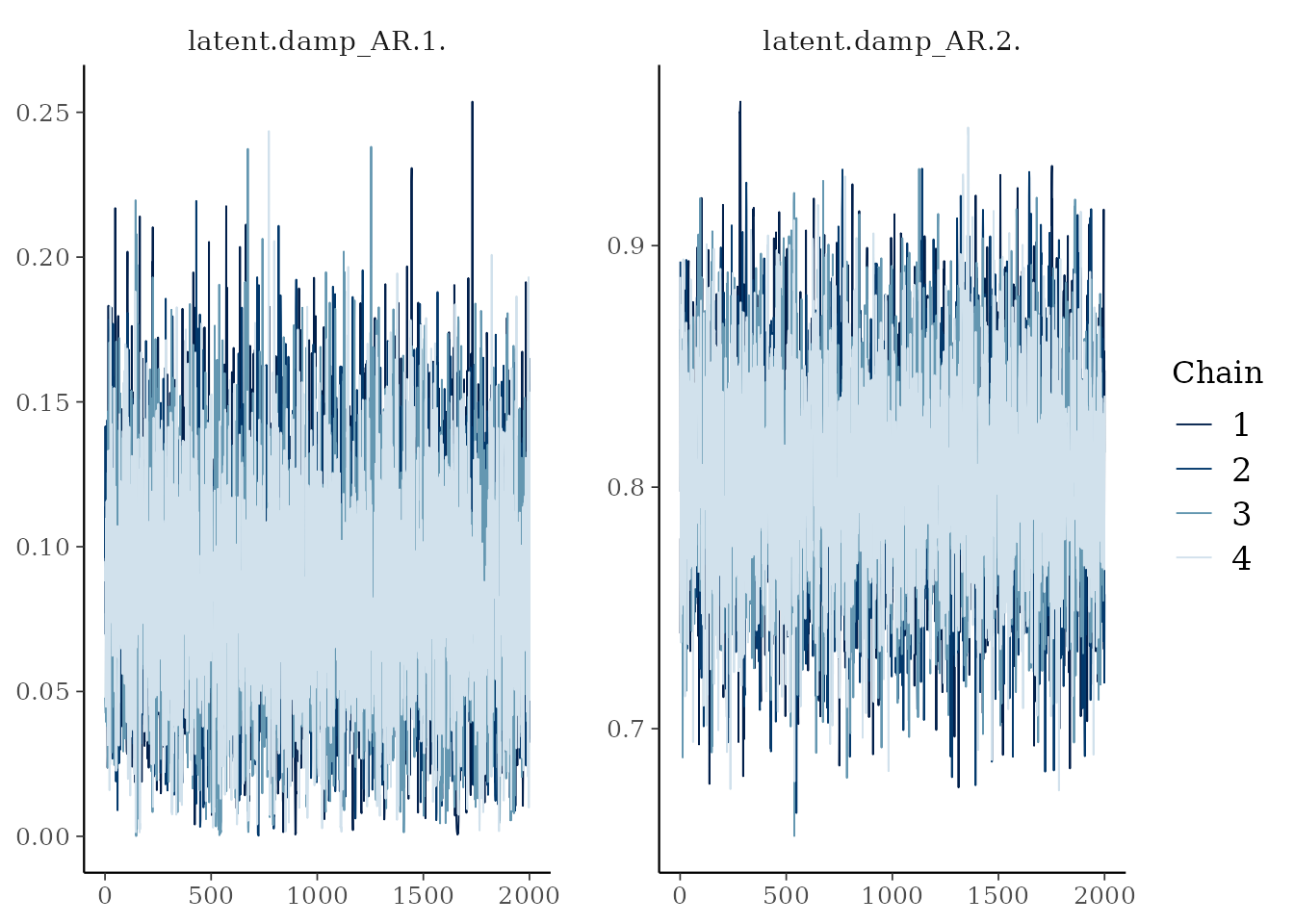
Model Comparisons
Comparing Latent Processes
Test sensitivity to AR order:
# AR(1) alternative - strong autocorrelation like AR(2)
ar1 <- AR(
order = 1,
damp_priors = list(truncnorm(0.8, 0.05, 0, 1)), # High autocorrelation
init_priors = list(norm(-1.0, 0.5)),
std_prior = halfnorm(0.5)
)
model_ar1 <- EpiProblem(
epi_model = renewal,
latent_model = ar1,
observation_model = negbin,
tspan = c(45, 80)
)
results_ar1 <- fit(model_ar1, data = full_data)
#> Generating Turing.jl model...
#> Running NUTS sampling...
#> Chains: 4
#> Warmup: 1000
#> Draws: 1000
#> Running Pathfinder initialization...
#> Pathfinder init failed, using default initialization...
#> Processing results...Compare the estimates from both models:
library(ggplot2)
library(patchwork)
p_ar2 <- plot(results, type = "Rt") +
ggtitle("AR(2) Model") +
coord_cartesian(ylim = c(0, 6))
p_ar1 <- plot(results_ar1, type = "Rt") +
ggtitle("AR(1) Model") +
coord_cartesian(ylim = c(0, 6))
p_ar2 + p_ar1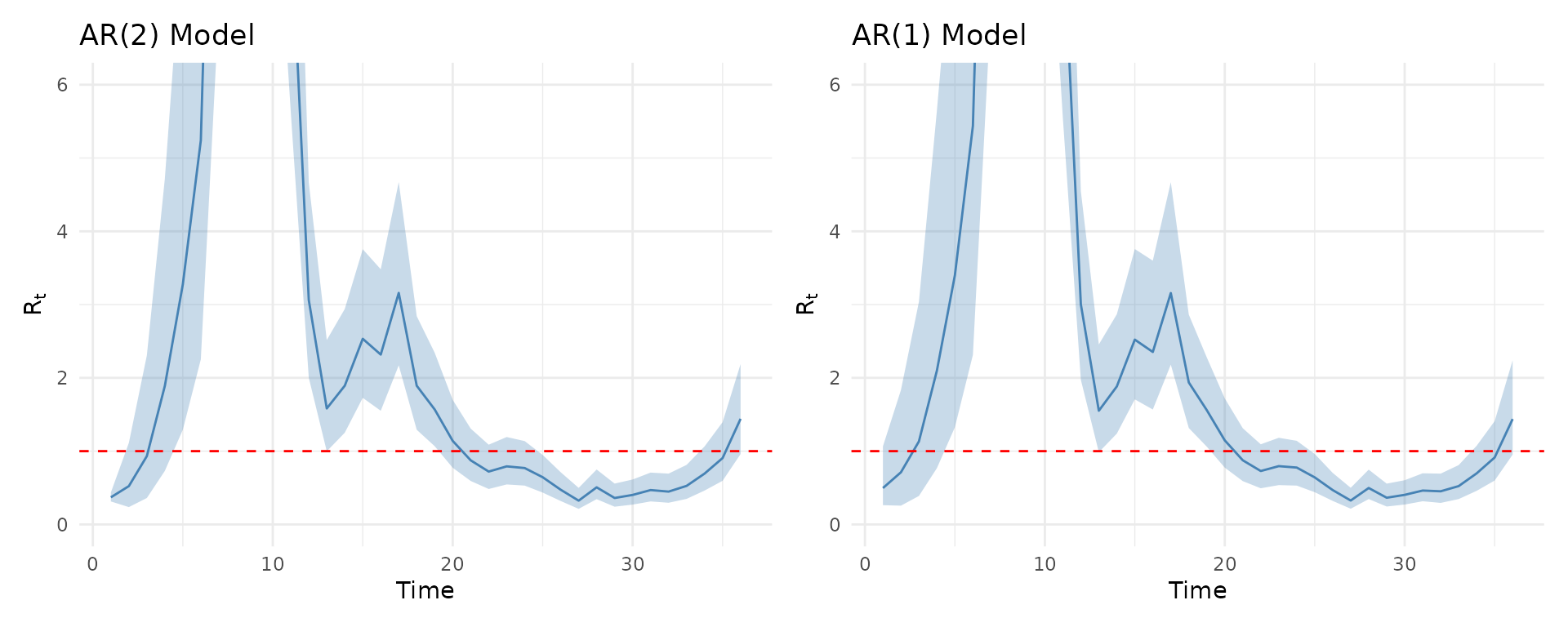
The AR(2) model typically produces smoother trajectories due to the additional lag term, while AR(1) may show more rapid fluctuations.
Adding Observation Delays
Account for incubation and reporting delays:
# Incubation period (~5 days)
obs_incubation <- LatentDelay(
model = negbin,
delay_distribution = lognorm(1.6, 0.42)
)
# Reporting delay (~2 days)
obs_full <- LatentDelay(
model = obs_incubation,
delay_distribution = lognorm(0.58, 0.47)
)
model_delays <- EpiProblem(
epi_model = renewal,
latent_model = ar2,
observation_model = obs_full,
tspan = c(45, 80)
)
results_delays <- fit(model_delays, data = full_data)
#> Generating Turing.jl model...
#> Running NUTS sampling...
#> Chains: 4
#> Warmup: 1000
#> Draws: 1000
#> Running Pathfinder initialization...
#> Pathfinder init failed, using default initialization...
#> Processing results...Compare models with and without observation delays:
p_no_delay <- plot(results, type = "Rt") +
ggtitle("Without Delays") +
coord_cartesian(ylim = c(0, 6))
p_delay <- plot(results_delays, type = "Rt") +
ggtitle("With Incubation + Reporting Delays") +
coord_cartesian(ylim = c(0, 6))
p_no_delay + p_delay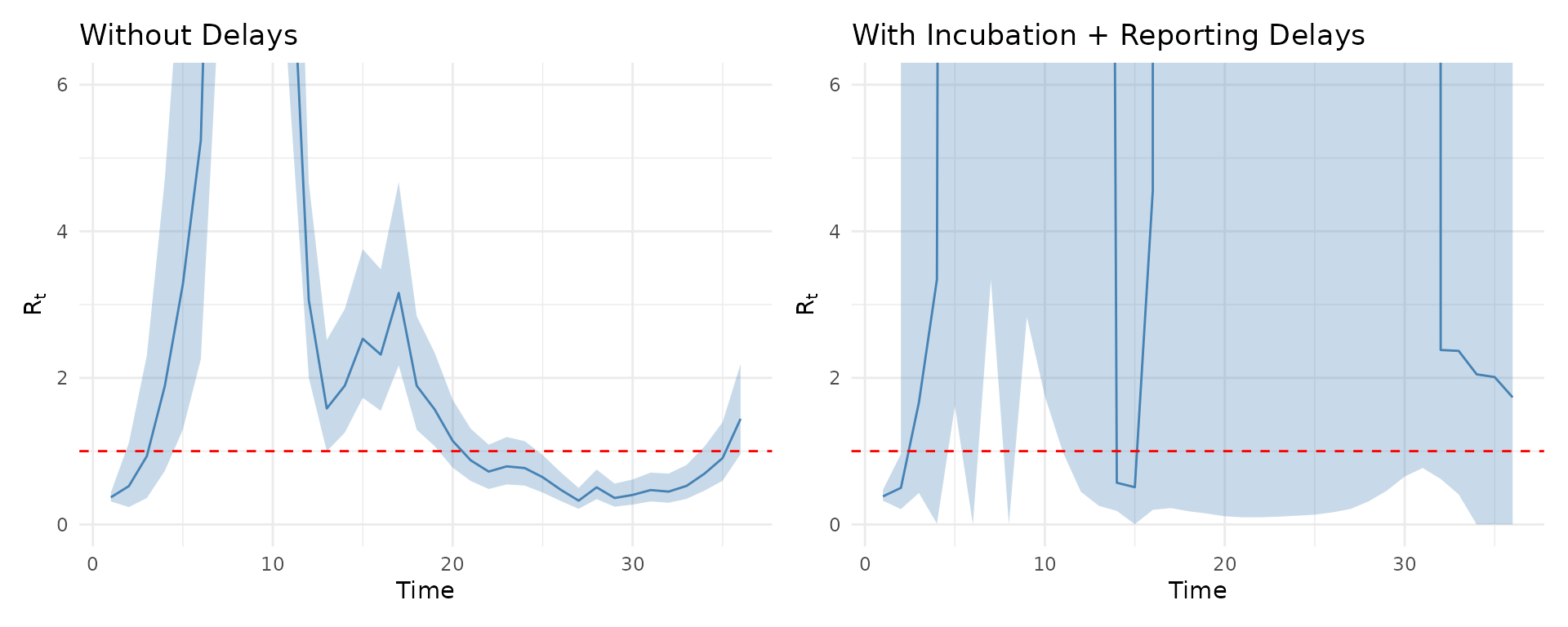
Accounting for delays shifts the trajectory earlier in time, as infections precede observed cases.
Interpretation
The Mishra et al. (2020) analysis demonstrated:
- Flexible estimation: AR(2) captures both smooth trends and rapid changes
- Uncertainty quantification: Bayesian credible intervals reflect parameter uncertainty
- Intervention detection: Sharp decline in coincides with public health measures
- Compositional flexibility: Easy to test alternative assumptions (AR order, delays)
Extensions
Try modifying the analysis:
- Different priors: Test sensitivity to prior choices
- Alternative latent models: MA, random walk, spline models
- Stratification: Separate models by age group or region
- Forecast evaluation: Hold-out validation of predictive performance
References
Mishra, S., Berah, T., Mellan, T. A., et al. (2020). On the derivation of the renewal equation from an age-dependent branching process: an epidemic modelling perspective. arXiv preprint arXiv:2006.16487.
Session Info
sessionInfo()
#> R version 4.5.3 (2026-03-11)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C
#> [3] LC_TIME=C.UTF-8 LC_COLLATE=C.UTF-8
#> [5] LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C.UTF-8
#> [9] LC_ADDRESS=C.UTF-8 LC_TELEPHONE=C.UTF-8
#> [11] LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C.UTF-8
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] patchwork_1.3.2 ggplot2_4.0.2 bayesplot_1.15.0
#> [4] posterior_1.7.0 EpiAwareR_0.2.0.9000
#>
#> loaded via a namespace (and not attached):
#> [1] tensorA_0.36.2.1 sass_0.4.10 utf8_1.2.6
#> [4] generics_0.1.4 stringi_1.8.7 digest_0.6.39
#> [7] magrittr_2.0.5 evaluate_1.0.5 grid_4.5.3
#> [10] RColorBrewer_1.1-3 fastmap_1.2.0 plyr_1.8.9
#> [13] jsonlite_2.0.0 backports_1.5.1 scales_1.4.0
#> [16] textshaping_1.0.5 jquerylib_0.1.4 abind_1.4-8
#> [19] cli_3.6.6 rlang_1.2.0 withr_3.0.2
#> [22] cachem_1.1.0 yaml_2.3.12 tools_4.5.3
#> [25] reshape2_1.4.5 checkmate_2.3.4 dplyr_1.2.1
#> [28] JuliaCall_0.17.6 vctrs_0.7.3 R6_2.6.1
#> [31] ggridges_0.5.7 matrixStats_1.5.0 lifecycle_1.0.5
#> [34] stringr_1.6.0 fs_2.0.1 ragg_1.5.2
#> [37] pkgconfig_2.0.3 desc_1.4.3 pkgdown_2.2.0
#> [40] pillar_1.11.1 bslib_0.10.0 gtable_0.3.6
#> [43] glue_1.8.0 Rcpp_1.1.1 systemfonts_1.3.2
#> [46] xfun_0.57 tibble_3.3.1 tidyselect_1.2.1
#> [49] knitr_1.51 farver_2.1.2 htmltools_0.5.9
#> [52] labeling_0.4.3 rmarkdown_2.31 compiler_4.5.3
#> [55] S7_0.2.1 distributional_0.7.0